Beyond Retinal: Machine Learning Models for Photochemical Control in Rhodopsins
Hector RCD Awardee Prof. Dr. Carolin Müller
Hector Fellow Prof. Dr. Klaus Robert Müller
Hector Fellow Prof. Dr. Peter Hegemann
The project is developing a machine learning framework to accurately predict the excited states of rhodopsins. To this end, a dataset of quantum chemical calculations on retinal derivatives in protein-like environments is being compiled and used for model training. The models are validated and refined through the repeated synthesis and spectroscopic analysis of specifically designed rhodopsin variants. The goal is to create a data-driven platform for the rational design of light-sensitive proteins and the accelerated development of new photoreceptors.
Rhodopsins are light-sensitive proteins that contain a covalently bound retinal chromophore as their photoactive unit. Although all rhodopsins share this common core, they exhibit a wide range of photochemical reactions—from simple E/Z isomerizations to multistep pathways—and thus perform different functions. This diversity arises from the delicate interplay between the intrinsic reactivity of the chromophore and the modulating effect of the surrounding protein matrix on the dark, excited, and photoproduct states.
A deep understanding of these interactions is essential for deciphering the molecular principles of biological light perception and for specifically controlling photochemical reactivity. Experimental methods such as time-resolved UV/Vis and Raman spectroscopy provide valuable data, but the ultrafast dynamic processes complicate their interpretation and often lead to speculative structure-property relationships. Quantum-theoretical simulations of the excited state offer mechanistic insights, but are practically inaccessible for the large chromophore-protein complexes of rhodopsins.
This project addresses this limitation by developing a machine learning (ML) framework that describes excited states in covalently bonded systems and uses rhodopsins as a model. The project brings together the expertise of Prof. Dr. Klaus Robert Müller (machine learning for chemistry and physics), Prof. Dr. Carolin Müller (high-quality QM/MM data and extension of ML models for excited states), and Prof. Dr. Peter Hegemann (synthetic, expressed, and spectroscopically characterized rhodopsin derivatives). The combination of mass-selective ion soft landing and ESR STM represents a groundbreaking methodological advancement that provides a modular platform for the controlled assembly of arbitrary molecular building blocks and their spin coupling, and can be seamlessly extended to larger biomolecules (e.g., metal proteins). In the long term, an open toolset will be created for the scientific community that links elementary surface physics with quantum information and sensor technology, laying the foundation for the next generation of molecular quantum simulators and optogenetic tools.
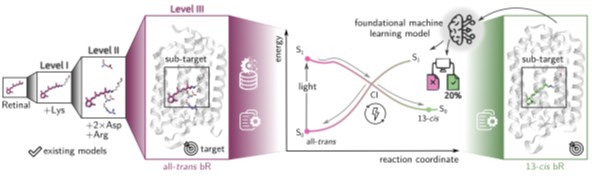
Figure 2: Illustration of the overarching project objective: Developing excited-state machine learning (ML) models to go beyond retinal model systems (left) to predict photoinduced phenomena of retinal within its native protein environment (colored boxes). This will be addressed by combining computational chemistry, machine learning, and spectroscopy to establish a foundational ML framework.
Supervised by

Carolin Müller
Chemistry, Informatics Hector RCD Awardee since 2024
Hector RCD Awardee since 2024
Klaus-Robert Müller
Informatics, Mathematics & PhysicsHector Fellow since 2023

Peter Hegemann
Biology, Chemistry & MedicineHector Fellow since 2015