PRECISE-CRC: PREvention through Causal Inference and Stratified Embeddings in ColoRectal Cancer
Hector Fellow Bernhard Schölkopf
Hector RCD Awardee Carolin Schneider
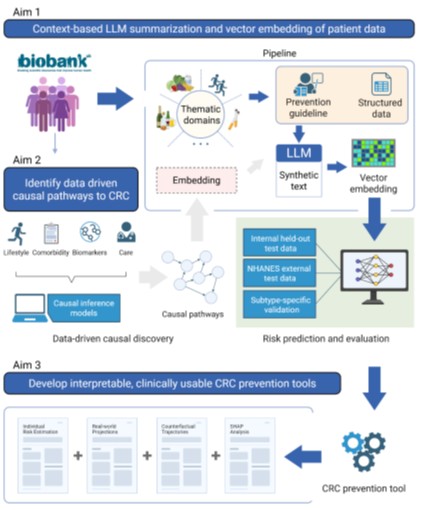
PRECISE‑CRC develops an innovative embedding framework that converts heterogeneous lifestyle, comorbidity, and care data from biobanks into structured patient summaries using a context‑aware large language model (LLM) and then transforms these summaries into high‑dimensional vectors with a text‑embedding model. Scalable causal methods (NOTEARS, Invariant Causal Prediction, Doubly Robust Estimator) are applied to identify true cause‑effect relationships for colorectal cancer and to calibrate the embeddings against objective biomarkers (omega‑3 fatty acids, BCAA, accelerometer‑derived activity, metabolomics). The resulting information is integrated into a digital prevention dashboard that provides individualized risk profiles, counterfactual scenarios and SHAP‑based explanations.
PRECISE‑CRC develops an innovative, embedding‑based analytical framework that converts the heterogeneous variety of lifestyle, comorbidity, and care data from large biobanks into a unified, semantically interpretable format. The core of the approach is a context‑sensitive large‑language model (LLM) that creates patient‑specific text summaries from tabular questionnaire and clinical information. These narratives are then transformed with modern text‑embedding models into high‑dimensional vector representations (embeddings) that compactly capture the complex relationships among dietary habits, physical activity, metabolic profiles, and disease status.
To move from mere associations to true causality, the embeddings are linked in a second analytical layer with advanced causal machine‑learning methods. Various scalable techniques—including NOTEARS, DoPFN, amortised inference for causal‑structure learning, Invariant Causal Prediction (ICP) and the Doubly Robust Estimator (DRE)—enable the construction of directed acyclic graphs that identify potential cause‑effect links between lifestyle dimensions (e.g., alcohol consumption, red meat intake, sedentary behavior) and the occurrence of colorectal cancer (CRC). A central quality feature of the project is the calibration of the algorithmically derived embeddings against objective biomarkers. Measurements such as omega‑3 fatty acids, branched‑chain amino acids, accelerometer‑derived activity metrics and comprehensive metabolomic profiles serve as independent validation to ensure biological plausibility and to avoid over‑fitting to subjective self‑reports. Correlations between individual embedding dimensions and the biomarkers are quantified with linear and non‑linear regression models; any identified discrepancies are fed back into iterative optimisation of prompt designs and weighting schemes within the embedding pipeline. The complete methodology is ultimately integrated into an interactive prevention dashboard.
The initiative is led by Prof. Dr. Bernhard Schölkopf, a world‑renowned pioneer in causal machine learning, and coordinated clinically by Jun.-Prof. Dr. Carolin Schneider, whose expertise in cancer prevention, biomarker stratification and digital health bridges the methodological advances to patient‑centred application. By linking semantic harmonisation, advanced causal inference, objective biomarker calibration and a user‑friendly visualisation platform, PRECISE‑CRC establishes a new standard for preventive oncology. The concept can be readily extended to other tumour entities or chronic diseases, thereby creating a sustainable infrastructure for causal embedding models in healthcare.
Figure 1: Overview of the proposed project
Supervised by

Bernhard Schölkopf
Informatics, Physics & MathematicsHector Fellow since 2018

Carolin Schneider
MedicineHector RCD Awardee since 2024